A RESEARCHER’S PERSPECTIVE: What Genetics Is Telling Us About Substance Use Disorders
A RESEARCHER'S PERSPECTIVE - from Brain & Behavior Magazine, February 2024 issue
By Sandra Sanchez-Roige, Ph.D.
Department of Psychiatry, School of Medicine University of California, San Diego
2018 BBRF Young Investigator
What have studies of the genetics of substance-use disorder so far uncovered? How can we transform these discoveries into concrete actions, effective interventions, and bring hope to those in need?
Substance use disorders have been my focus throughout my career. They are among the most common psychiatric conditions. Causation is complex—embedded in a web of environmental and genetic factors. Prevention, diagnosis, and treatments remain limited.
Over the past 6 years, there has been an explosion of large studies aimed at finding the genetic factors that may be associated with substance use disorders. These studies are vital, because they can reveal biological mechanisms of substance use disorders that can be targeted for new interventions, and thus make a significant impact in people's lives. This fuels my research.
It has been inspiring to see the progress in human genetics. At the forefront have been genome-wide association studies, or GWAS. These studies are designed to scan across our human genome in search of genomic regions that may be associated with a trait like problematic drinking.
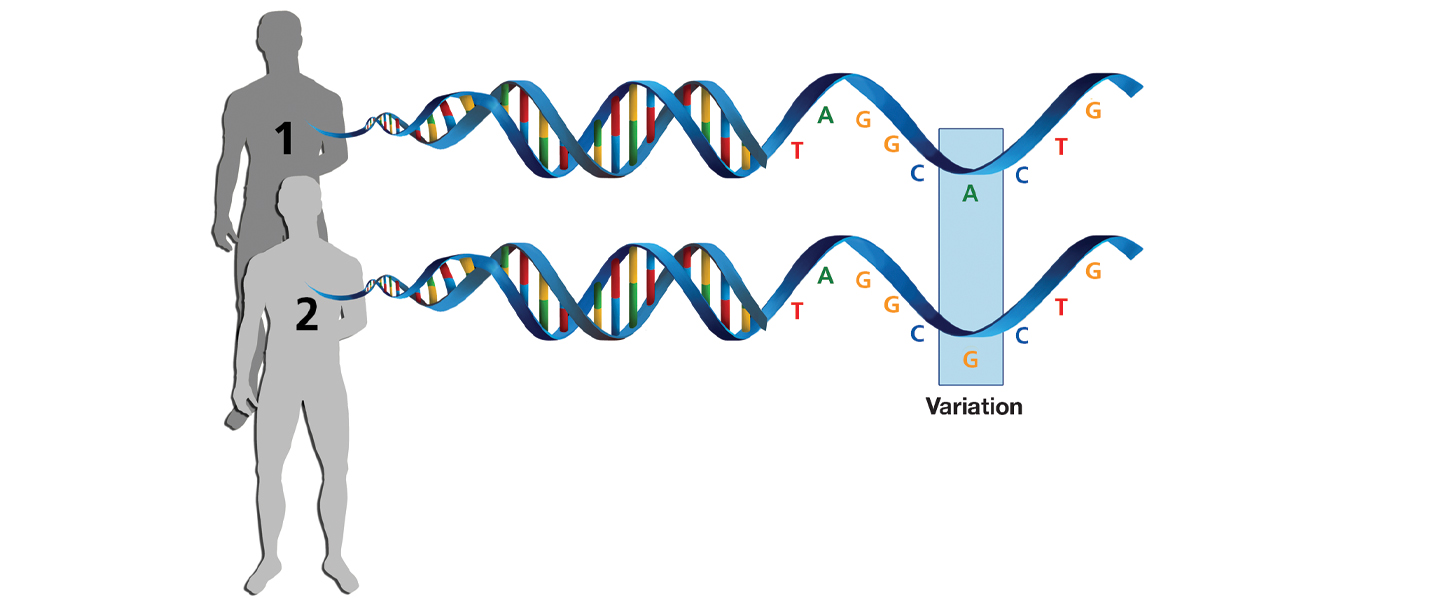
Your genome is the complete sequence of DNA that you have in all the cells in your body that is used as the blueprint to build all the parts of the body. A DNA strand is made up of a sequence of DNA “letters”—abbreviated as G, C, T, and A. Although much of the genome (around 3 billion pairs of DNA letters) is the same in all humans, some letters can be different between individuals. For example, at a particular spot in the genome, you might have an A, whereas someone else has a G.
An individual’s unique configuration of genetic variation across the genome is called their “genotype.” A GWAS measures millions of these genomic variants and correlates each one with the trait that is being studied. Alcohol dependence is an example of a trait we can study with GWAS. Scientists often call them “phenotypes,” but I am going to use the friendlier term “trait” in this article.
GWAS have proved to be extremely successful tools, but in order to obtain meaningful results, we’ve learned that we need extremely large sample sizes— hundreds of thousands of people whose genomes we have received permission to scan (anonymously, of course—a process called “de-identification”). We must also have data about an individual’s traits. For example, in order to conduct GWAS of alcohol dependence we must have a way to identify those who have an alcohol dependence diagnosis and contrast their genomes with those who do not. Computational methods to analyze the vast amounts of data from these huge cohorts have become more refined over the years.
How do we measure substance use disorders? Doctors can very accurately measure our blood pressure; they have very precise instruments for that. But when we go to a psychiatrist’s or a psychologist's office, they will ask us a series of questions to determine whether we meet certain criteria for a disorder diagnosis. And if we carry two or more of the symptoms outlined in the DSM-V diagnostic manual, we would be diagnosed with a substance use disorder.
Using the diagnosis as defined in the DSM, we could recruit lots of people with or without a diagnosis of alcohol use disorder and perform a GWAS. I have been part of major efforts of this kind led by the largest international consortium on psychiatric genetics, the Substance Use Disorder Workgroup of the Psychiatric Genomics Consortium. One of the landmark papers by this group, published in 2018, focused on assembling multiple cohorts to get a sample large enough to perform a statistically meaningful GWAS of alcohol dependence. In this study, our subjects were identified according to the diagnostic criteria of the DSM manual. After much rigor and effort, this study was able to reveal only one locus—one location in the human genome—with a statistically significant association with alcohol use disorder. This area contains one of the genes that regulates how ethanol is metabolized in the body. We realized that much larger sample sizes would be needed to achieve the statistical power to find other risk locations in the genome related to this particular trait.
HOW TO GET BIGGER AND BETTER SAMPLES
Over time, research began to reveal that substance use disorders, like all complex traits, are highly polygenic. This means that many genetic risk variants—hundreds or even thousands—are involved in vulnerability for the trait. Commonly occurring risk variants are thought to each have a very small impact on total risk for the trait (here, alcohol dependence). Yet we realized that in our GWAS cohorts we may be including high levels of heterogeneity—differences between people who display the trait we are looking at. For example, there are over 2,000 unique combinations of the DSM diagnostic criteria that can result in a substance use disorder diagnosis. That means two people can be given the same diagnosis, yet exhibit the substance use disorder in very different ways.
To complicate things even further, substance use disorders develop over time, starting with experimental use, leading to regular use, compulsive use, then, often, cessation, and also, often, relapse. It is possible that different genes may impact or have a different role at different stages of substance use disorders. If we’re not careful in assembling our study populations, we may inadvertently obscure signals from the genome that may be relevant to each of these stages, signals which could potentially have enormous therapeutic value.
Back in 2016, I joined Dr. Abraham's Palmer laboratory as a postdoc. Dr. Palmer is a 2006 and 2003 BBRF Young Investigator. The challenge we wanted to solve was related to the difficulties of putting together samples to perform GWAS. We asked, “where can we get high volumes of good data that may be relevant to substance use disorders?” The answer was: with a consumer genetics company, 23andMe, Inc. The beauty of working with them is that they have already genotyped millions of people.
We paid 23andMe to deploy an online survey capturing different aspects of substance use that we compiled with input from psychologists. 23andMe users who participate in the company’s genome research efforts were asked to fill out the survey. This was completely voluntary with informed consent.
This survey included a 10-item questionnaire, called the Alcohol Use Disorder Identification Test (or “AUDIT”) that measures past-year alcohol use. We collected 25,000 AUDIT responses from 23andMe research participants. We then combined this data with additional AUDIT data from another population- based cohort, the UK Biobank, which has genotype and trait data for half a million participants.
We aggregated the data from these two datasets and performed a GWAS, which uncovered 10 genome locations associated with elevated risk for problematic alcohol use, nine more than the inaugural 2018 GWAS of alcohol dependence that I mentioned. It was reassuring to learn that among the 10 risk locations identified in this GWAS, one was the location that we identified previously, which included the gene for an enzyme that metabolizes ethanol.
The beauty of the AUDIT questionnaire is that it can distinguish alcohol use from misuse. For example, the first three items measure aspects of consumption, such as the frequency of drinking, quantity of drinking, and patterns of binge drinking (drinking too much over a short time period). We called this version of the questionnaire AUDIT-C. Another variation measuring problematic consequences of alcohol use, such as causing injury to oneself or others as a consequence of drinking, we named AUDIT-P.
We performed two separate GWAS based on responses to AUDIT-C and AUDIT-P. It was clear that some genome risk locations, or “loci,” consistently appeared (for example, the ethanol-metabolizing enzyme genes I’ve mentioned, which are on chromosome 4). But we also saw that the overlap was incomplete, suggesting that the genetic architecture of alcohol use is not the same as the genetic architecture of alcohol misuse. This is important; it calls attention to the need to distinguish these two core aspects relevant to alcohol use disorder.
How closely related were these various AUDIT traits to clinically defined alcohol dependence (i.e., based on the DSM criteria)? We used a revolutionary statistical method to perform genetic correlations, which allowed us to estimate the genetic factors shared between two traits. The uniqueness of this method is that unlike “trait” correlations, which are performed using the same individuals, genetic correlations can be performed across pairs of traits that are measured in independent cohorts.
We performed genetic correlations between AUDIT-C and AUDIT-P and alcohol dependence, and we identified very strong, significant genetic correlations between the traits. These findings, to my view, are extremely important, because they illustrate that we could combine clinical traits and reach unprecedented sample sizes. Thanks to support from BBRF, Dr. Hang Zhou, a 2018 BBRF Young Investigator, combined clinical data with AUDIT data, revealing 110 risk locations in the genome associated with problematic alcohol use, a dramatic increase from our original study back in 2018.
So, to sum up so far: Using AUDIT, we have shown that alcohol use and misuse have a different genetic basis, and we have uncovered hundreds of novel genes associated with problematic alcohol use. I've also told you that we've begun to stiudy the genetic underpinnings of the different addiction stages.
We have also learned that alcohol use disorders, like all complex conditions, are not single-gene disorders and that hundreds to thousands of commonly occurring genetic variants, each likely with small impact on total risk, are involved in the condition.
“HAVE YOU EVER MISUSED AN OPIOID?”
Another study I’d like to discuss pertains to aspects of prescription opioid use. The metrics for the current opioid epidemic in the U.S. are shocking. Almost 130 people die every day from opioid overdose. The majority of opioid users initiate with prescription pain relievers. Because prescription opioids are widespread in medical settings, we believe that prescription opioid misuse is be a trait that could be captured in, for example, the 23andMe population.
In 2018, we extended our 23andMe survey to 125,000 people, and one of the added questions pertained to taking prescription painkillers not as prescribed. To our surprise, 20% of the 23andMe research participants reported having at least once taken opioids not as prescribed. We wondered if a trait like this could generate a genetic signature that is highly correlated with opioid use disorder.
We performed a GWAS and identified two significant risk loci. The strongest signal was with variants in the gene KDM4A, which, intriguingly, was recently associated with opioid use disorder as diagnosed by clinicians in an independent study.
We found that the KDM4A gene interacted with multiple drugs, including serotonin reuptake inhibitors (SSRI antidepressants) as well as disulfiram (used to treat chronic alcoholism), medicines often prescribed for disorders known to co-occur with opioid use disorder. We also found interaction with drugs affecting the dopamine system that are known to influence neural circuits associated with reward and reinforcement—which are critical in substance use disorders.
Even more important, the trait of opioid prescription misuse shows strong genetic correlations with results of the largest available GWAS of opioid use disorders. We also identified strong genetic correlations with over 200 other outcomes, particularly other substance-use traits, pain, pain medications, as well as associations with risky behaviors.
We were concerned that maybe we were just picking up on a signal that had little to do with opioid misuse, but more to do with risky behaviors—people who may be inclined to misuse opioids may also be inclined to risky behavior in general. We used statistical tools that suggest the signal we captured is primarily specific to opioids and not merely risk- taking.
So here again, by asking a single simple question in a cost-effective way—as we did when we asked 23andMe research participants about whether they had ever misused an opioid—we were able to help discover something of broad importance about the genetic basis of opioid use disorders.
Encouraged by these findings, my lab, in collaboration with Dr. Palmer and 23andMe, have launched what we call the Prescription Opioid Genetics Study in a cohort of half a million people from multiple ancestries. This survey has already been deployed and we are in the process of collecting data. All subjects in the cohort have a history of using prescription opioids at least once in their life. We will incorporate data on pain, trauma, and other conditions that are known to intersect with opioid use.
We hope this will further our understanding of opioid use disorder as well as our understanding of the intricate relationships between prescription opioid use and misuse and the relationship with mental and physical health. We hope to finish assembling this data set in 2024.
HOW IMPULSIVITY CONTRIBUTES TO SUBSTANCE MISUSE
Let’s turn now to another trait, impulsivity: thoughts or actions that are poorly conceived, prematurely expressed, and that often result in undesirable consequences.
Many neuropsychiatric disorders are associated with impulsivity, including substance use disorders. Impulsivity is involved at multiple stages of vulnerability for substance use disorder, including the transition from regular to harmful use, as well as in relapse. By studying the fundamental nature of this trait of impulsivity, which is present in each one of us at a higher or lower level, it is our hope to learn more about the biology underpinning multiple conditions characterized by excessive impulsivity levels, including substance use disorders.
As part of our collaboration with 23andMe, our survey included several well-established questions that capture different facets of impulsivity. For example, we asked respondents to assess themselves on the statement: "I quite enjoy taking risks." We also asked people to respond to the statement: "I do things without thinking."
We collected 150,000 responses on eight impulsive personality traits and we performed independent GWAS studies of each of these traits. We learned that impulsivity is heritable, which means that the extent to which we are more or less impulsive, or the way that we responded to the previous questions, can be attributed, in part, to genetic factors. We found a heritability of 10% for impulsivity.
This means that only about 10% of impulsivity can be explained by genetic factors. And this suggests that even though we, as geneticists, are interested in finding biological causes that contribute to behavior, the environment plays an equally or even more important role in shaping how we feel and how we act.
We also learned that the impulsivity-related traits are genetically correlated, but that the overlap is incomplete, emphasizing what has been known in neuroscience for a long time: each impulsivity trait is governed by different biological mechanisms. These impulsivity traits are also genetically correlated with substance use traits, traits spanning nicotine dependence, and alcohol, cannabis, and opioid use disorders. The challenge for future studies is to disentangle the nature of this complex web of correlations.
The GWAS for the eight impulsivity traits we measured identified 16 genomic regions associated with impulsivity. The frustration all geneticists share is that GWAS are correlation studies and can only point to regions in the genome that are associated with a trait. The variations we find in these locations do not cause impulsivity [chart, facing page]. They don’t even indicate what we call the “directionality” of the association.
For example, even though we and others have robustly established that variations in the gene CADM2 are associated with impulsivity, we still have little understanding about the mechanisms in the brain through which this gene influences behavior. My BBRF Research Partners grant (supported by Families for Borderline Personality Disorder Research) enabled us to produce mice with the variation in the mouse version of this gene, called Cadm2. (Borderline Personality Disorder is also characterized, in part, by impulsive behavior). In the genome there are two copies of each gene, and in mice we can selectively remove or deactivate one or both genes to directly test how it impacts behavior. Using mice with manipulated Cadm2, we tested how this gene contributes to performance in a broad battery of behavioral tasks that included measures of impulsivity.
In one test that measures “risky” responses in mice, we found that mice lacking one functional copy of the Cadm2 gene showed less risky behavior. This is an indication, in one case, of the direction in which a specific genome variation affects a given trait—in this case impulsivity.
We also measured other facets of impulsivity. Loss of both functional copies of the Cadm2 gene in mice resulted in less impulsive responding to a task. Beyond impulsivity measures, we did not observe deficits in tasks we had the mice perform to measure anxiety-like behavior. This showed that this gene, Cadm2, at least in the ways we were able to assess in our study, seems to be specific to impulsivity and not general aspects of behavior.
TURNING SIGNALS OF RISK INTO BIOLOGICAL KNOWLEDGE
Our knowledge of the genetic underpinnings of traits such as impulsivity or problematic substance use are not meant to replace clinical diagnosis of substance use disorders. But they do allow us to dissect, in this case, substance use disorders; they provide a more granular biological understanding of them, which enables us to move toward translational research in years to come.
GWAS have been tremendously successful. And with the availability of newer, larger-scale data sets (one is called All of Us), or other academic initiatives such as the PsycheMERGE consortium (with access to longitudinal data from millions of individuals), it is almost certain that the number of risk loci associated with substance use disorders will continue to grow.
But: is increasing sample sizes for GWAS—with the hope of identifying more risk locations in the genome— all we should be doing? When we perform such studies, moreover, what are we going to do with all the newly identified risk loci in the genome? In other words, how are we going to turn these statistical signals of risk into biological knowledge?
If we can do this, it is quite possible that we will find better targets that could have treatment and prevention value. We are working now on developing new methods that go beyond identifying risk genes like CADM2. We seek to generate 3D high-resolution maps of where and when such genes impact operations of the brain.
We must increase the diversity of the population samples whose genomes form the basis for our research. Most of the studies that I have presented here are dominated by one group of genetically similar individuals, namely individuals of shared European ancestry. Some of us are working on translating and integrating research findings across ancestries to enable equitable health research and therapeutic innovations.
It is going to be very exciting to see the impact upon our knowledge of substance use disorders moving forward. Our ultimate goal is to translate some of the most promising findings to the clinic to provide relief to those who suffer from substance use disorders.
A final appeal to those struggling with this condition or to those who have a family member or a beloved friend who does. If you think, "How can I help genetic studies?" please know that it is very important that we increase community participation.
We encourage others to become involved. These studies are very costly, not only in terms of funding, but also because it is incredibly challenging to reach the large sample sizes that make our studies statistically robust and thus meaningful to the discovery of new therapies. We need help from the public.
As some of the work I’ve presented here suggests, we can learn a lot about substance use disorders by learning about people who do not, in fact, have substance use disorders. For example, there is much to be gleaned by studying how impulsive a person is, or how a person responded to opioids the first time they used one. There is a great deal we have already learned by asking questions like these of anyone who wants to help.
Written By Peter Tarr, Ph.D.
Click here to read the Brain & Behavior Magazine's February 2024 issue